Fully Sharded Data Parallel (FSDP)
With FSDP, not only do we shard the batch over multiple chips like with Data Parallelism, we also shard the optimizer state, the gradients and the parameters over multiple chips- This allows training models that are orders of magnitude larger than a single chip's memory.
Gather-Compute-Discard
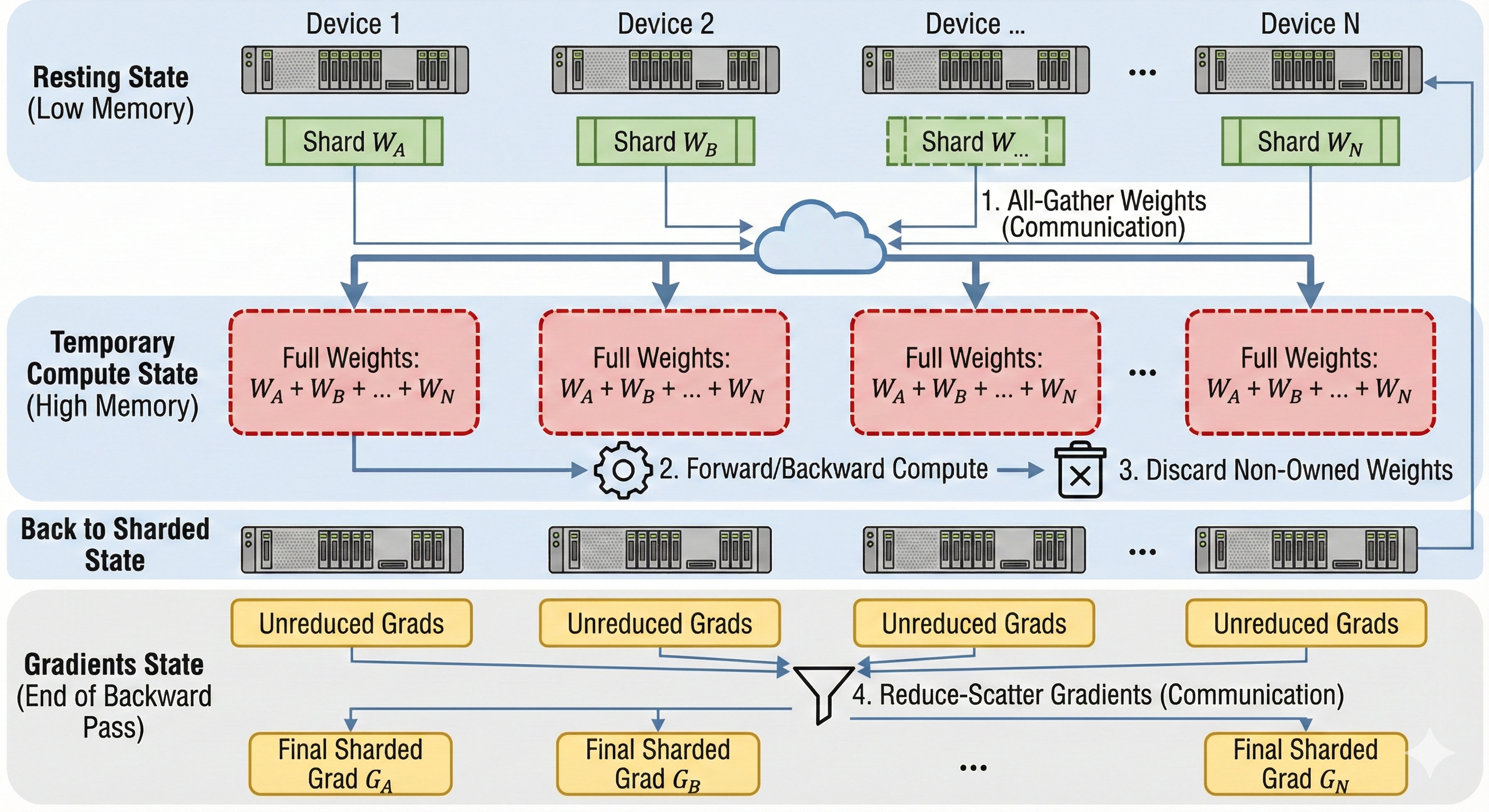
The main mechanism behind FSDP is called Gather-Compute-Discard. Since parameters are sharded, a device cannot compute a layer immediately. It must first "borrow" the missing data from its neighbors.
- Shard: We initially fully shard tensors to reduce the per-chip memory.
- All-Gather (Weights): Before the forward pass of a layer, we All-Gather the parameters so that each chip momentarily holds a full replica of that specific layer.
- Compute: We compute the forward/backward pass with the full layer.
- Discard (Weights): We delete the parts of the tensor our chip did not initially owned to reduce memory requirements.
- Reduce-Scatter (Gradients): After the backward pass, instead of All-Reducing (which keeps a full copy of gradients everywhere), we Reduce-Scatter the gradients. Each chip ends up with only the specific chunk of gradients corresponding to the parameters it owns.
Note: The communication is typically overlapped with computation to hide latency.
The three stages
Sharding more tensors means increasing the amount of All-Gathered data. Ideally we would shard as little as possible. Nonetheless, if only sharding the optimizer's state is not enough, we need to shard the gradients as well, or even the model parameters.
We often refer to the levels of sharding as the Three Stages of ZeRO, after Deepspeed's ZeRO paper.
| Stage | What is Sharded? | Memory Savings | Communication Overhead |
|---|---|---|---|
| ZeRO-1 | Optimizer States only | ~4x reduction. (Optimizer states are typically 75% of training memory). | Minimal (Same as DDP). |
| ZeRO-2 | Optimizer + Gradients | 8x reduction. | Minimal. |
| ZeRO-3 | Opt + Grads + Parameters | Linear reduction (\(1/N\)). Allows fitting massive models. | High. Requires All-Gather before every layer. |
Why is the optimizer state so large?
For every parameter, we hold:
- 2 bytes (bf16 weight)
- 2 bytes (bf16 gradient)
- 12 bytes (f32 optimizer state: master copy, momentum, variance)
Total: 16 bytes per parameter. Sharding just the optimizer states (12 bytes) removes 75% of the memory footprint without adding any extra communication steps (since All-Reduce and Reduce-Scatter transfer the same volume of data).
Pros and Cons
FSDP should only be used during training. It saves memory but doesn't speed up the math for a single sample; in fact, the communication overhead would make generation slower.
Code
Since our Fake API does not expose the optimizer's state, we will focus on sharding the model's weights and gradients.
To implement our initial unsharded model with FSDP, we need to change several things:
- Load a subset of the weights from our checkpoint.
- All-Gather the weights before each layer in the forward pass.
- Delete the gathered weights after using it (it would be implicit in an ML framework like Jax.)
- Reduce-Scatter the gradients after each layer of the backward pass.
Let's implement our 2 layers model with a ReLU activation, we shard the Model dimension across N devices such that each device holds Model/N:
- Correctness Note: We do not need to
Reduce-Scatterthe activations gradients, just the weights gradients. - Performance Note: In a production system, we would overlap the
All-Gatherfrom layerNwith the computations from layerN-1. We would also overlap theReduce-Scattersfrom layerN-1with the gradients computations from layerN.
class FSDP(ShardedEngine):
def __init__(self, model_dim: int, hidden_dim: int):
# How much data does a single device hold on the model axis
self.local_model_dim = model_dim // self.num_devices
self.w0 = np.zeros((self.local_model_dim, hidden_dim), dtype=np.float32)
self.w1 = np.zeros((hidden_dim, self.local_model_dim), dtype=np.float32)
# Context tape to store activations for backward pass
self.activations = []
def load_checkpoint(self, params: dict[str, npt.ArrayLike]) -> None:
# Indices in the global array that belong to this device
global_start_idx = self.device_id * self.local_model_dim
global_end_idx = global_start_idx + self.local_model_dim
# Load weights into local memory
self.w0[...] = params['layer_0/weights'][global_start_idx:global_end_idx, :]
self.w1[...] = params['layer_1/weights'][:, global_start_idx:global_end_idx]
def forward(self, x: npt.ArrayLike) -> npt.ArrayLike:
# -- Same as in single device --
self.activations.append(x)
# All-Gather w0
w0_global = self.all_gather(self.w0, axis=0)
# -- Same as in single device --
z = np.einsum('bd,df->bf', x, w0_global)
# Delete gathered data
del w0_global
# All-Gather w1
w1_global = self.all_gather(self.w1, axis=1)
# -- Same as in single device --
self.activations.append(z)
x = relu(z)
out = np.einsum('bf,fd->bd', x, w1_global)
# Delete gathered data
del w1_global
return out
def backward(self, grads: npt.ArrayLike) -> dict[str, npt.ArrayLike]:
"""
grads: Incoming gradient dL/d(Output) of shape (Batch, Model_Dim)
"""
# -- Same as in single device --
z = self.activations.pop()
h_relu = relu(z)
w1_grad = np.einsum('bf,bd->fd', h_relu, grads)
# Reduce-Scatter w1_grad
w1_grad = self.reduce_scatter(w1_grad, op='avg', axis=1)
# All-Gather w1 again
w1_global = self.all_gather(self.w1, axis=1)
# -- Same as in single device --
grads = np.einsum('bd,fd->bf', grads, w1_global)
# Delete gathered weights
del w1_global
# -- Same as in single device --
grads = grads * (z > 0)
x_input = self.activations.pop()
w0_grad = np.einsum('bd,bf->df', x_input, grads)
# Reduce-Scatter w0_grad
w0_grad = self.reduce_scatter(w0_grad, op='avg', axis=0)
return {'layer_0/weights': w0_grad, 'layer_1/weights': w1_grad}